Using Vish for Cactus/Carpet
Installing Vish
Vish runs under Windows XP/32bit, Windows Vista/32bit, Linux 32/64 bit, Mac OS X. Windows 64 bit is under investigation.
The preferred mode to get Vish is by compiling from source.
For convenience, binary installers are provided via the
CCT Vish Project Page,
it is updated occasionally and might not be the most recent version.
This CCT Vish Project Page contains all online available
information about Vish anyway, just its webdesign is not optimal (volunteers welcome).
Source code access is provided via SVN at the
Origo development server
(a swiss version of sourgeforge).
You will need to register for an account at Origo and
send an email to werner(at)cct.lsu.edu .
Eventually we might also provide Linux binaries for convenience
if this turns out to be preferable.
Compiling Vish is (at least supposed to be) easy.
There are very few dependencies on external libraries.
The biggest external package is Qt. This needs to be installed
first for the user interface (not for the Vish kernel, which is
completely independent from the graphical user interface - it
will compile fine, but once the GUI is to be compiled it will
lead to errrors if Qt is not available). The only other
external dependencies are OpenGL (not unusual for visualization
software) and freetype (but only used for a plugin to write
3D captions - Vish will run without).
More details at the Vish Development documentation at Origo.
Preparing Cactus/Carpet data for Vish
Vish reads HDF5 files in special layout, called the "F5" layout, a
shortcut for "Fiberbundle HDF5"
(see its dedicated page for more detailed information).
The HDF5 files produced by the standard cactus I/O thorns cannot be read
by Vish, they need to be converted into F5 layout.
A couple of converters are provided in the fish/fiber/F5apps/converters/
subdirectory that comes with the Vish SVN:
- CCTKtoF5 - for Cactus unigrid output (PUGH driver)
- CARPETtoF5 - for Carpet AMR output (Carpet driver)
- CarpetInterpToF5 - for Carpet interpolated arrays
- MULTIPATCHtoF5 - for Carpet multipatch output
To compile the respective converter, change into the respective subdirectory
and type "make". The executable will reside in the bin/ subdirectory of the
Vish installation.
Each converter should tell its required command line arguments when invoked
without arguments. In general it will have some intelligence built in to
convert a set of cactus output files into one single .f5 file. Note that
it is not a conversion from "one" cactus .h5 file to one .f5 file,
but the information contained in one .f5 file will encompass many .h5 files.
To inspect the contents of the resulting .f5 file use the
tool F5/apps/tools/F5ls (same installation instructions as with the
converters). It operates similar to the "h5ls" tool, but provides information
about the file content on a semantically higher level.
Alternative to aftermath data conversion, there exist Cactus and Carpet
thorns that can produce F5 files natively during the simulation run.
They are available through the F5 Origo Development Page.
Visualizing Cactus Data in Vish
General User Interface Concepts
Once you have a .f5 file, start Vish with this file given on the
command line (or in Windows, by clicking on it). The vish executable
will be in bin/vish in the vish installation directory.
Assume your file is called MergingNeutronStars.f5,
as it is available from our Scientific Visualization Data Page.
It should look like this:
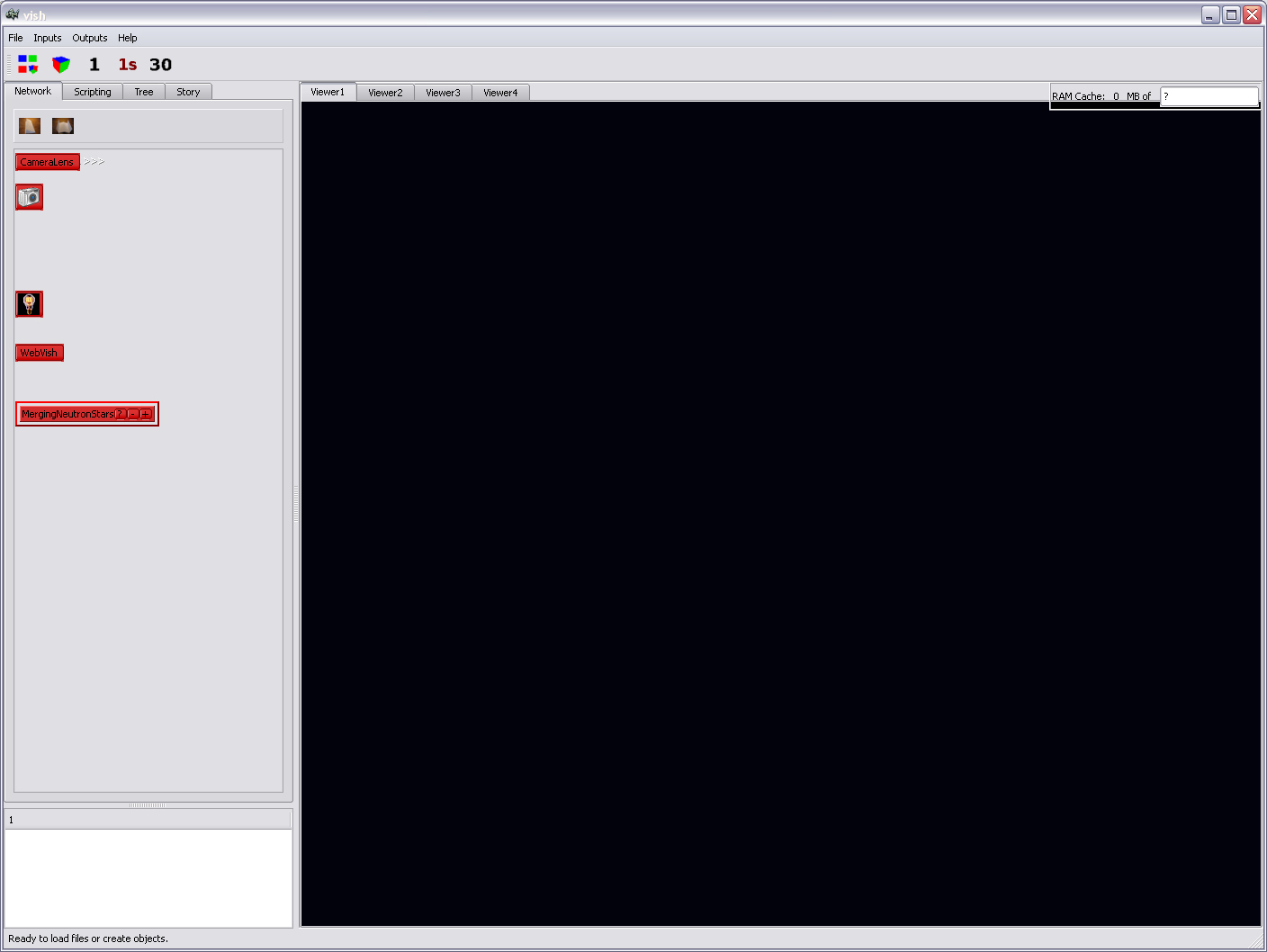
Note the red icon in the "Network" tab on the left side with the same
name as the filename, "MergingNeutronStars".
Rightclick on this icon, find a menu popping up. Go the "Output"
submenu, and from there to the "Fiber" submenu, where to select
the "Grid" submenu. In the following text, we will refer to
an operation of this kind as selecting the "Fiber/Grid" output.
You will get a new, orange icon:
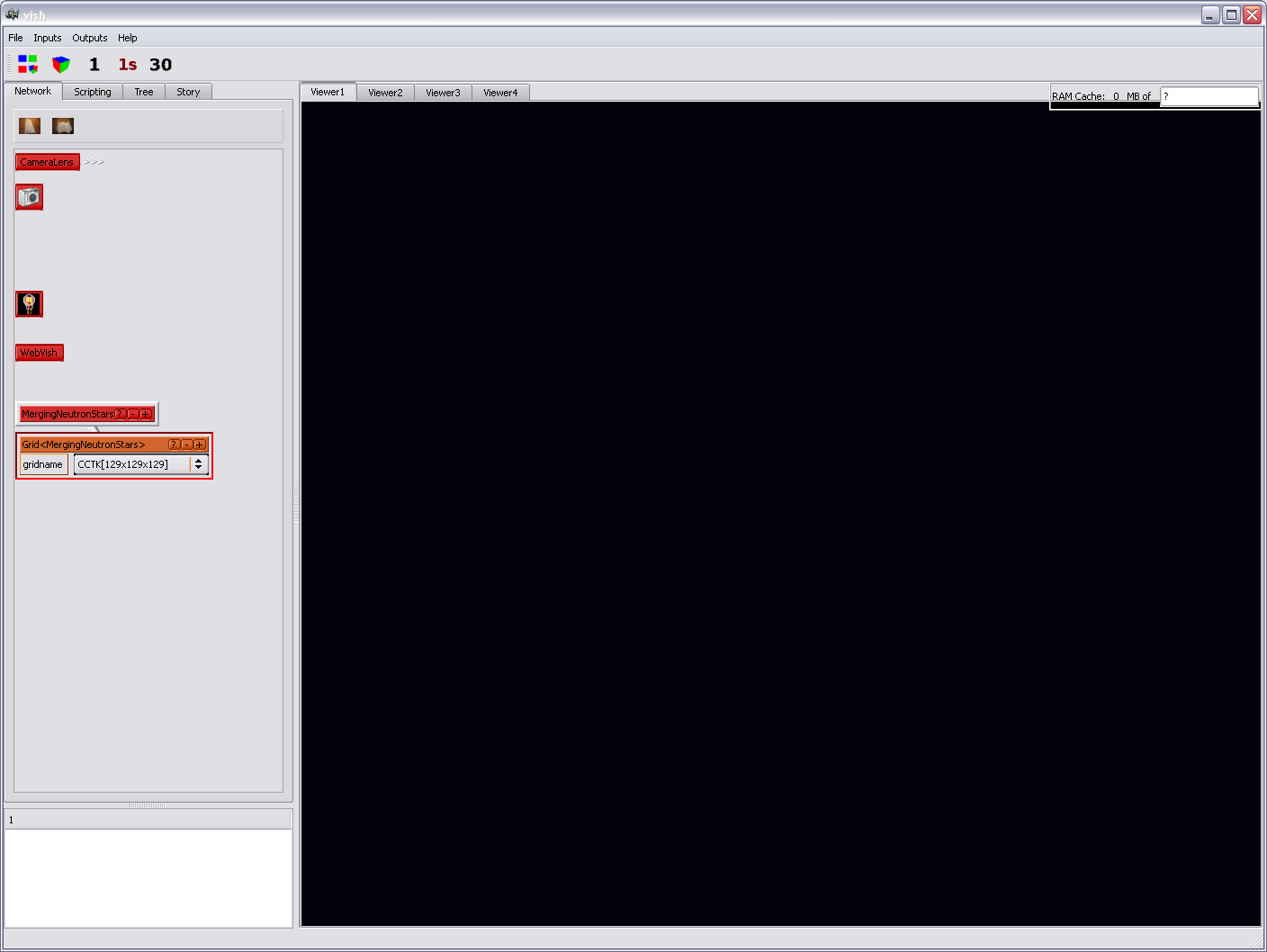
The orange icon will contain one selection element which in case of
the said data set says "CCTK[129x129x129]". When converting data
from cactus output, there will only be one such selection element
in the current scheme. There might be more if .f5 files from
different cactus runs are merged, but such a merging tool has
not yet been implemented.
On the orange icon (we call this the Grid object), add an
Display/BoundingBox object (from the Output context menu, as
described above. You will get a new yellow-orange icon, and
an orange icon "Time":
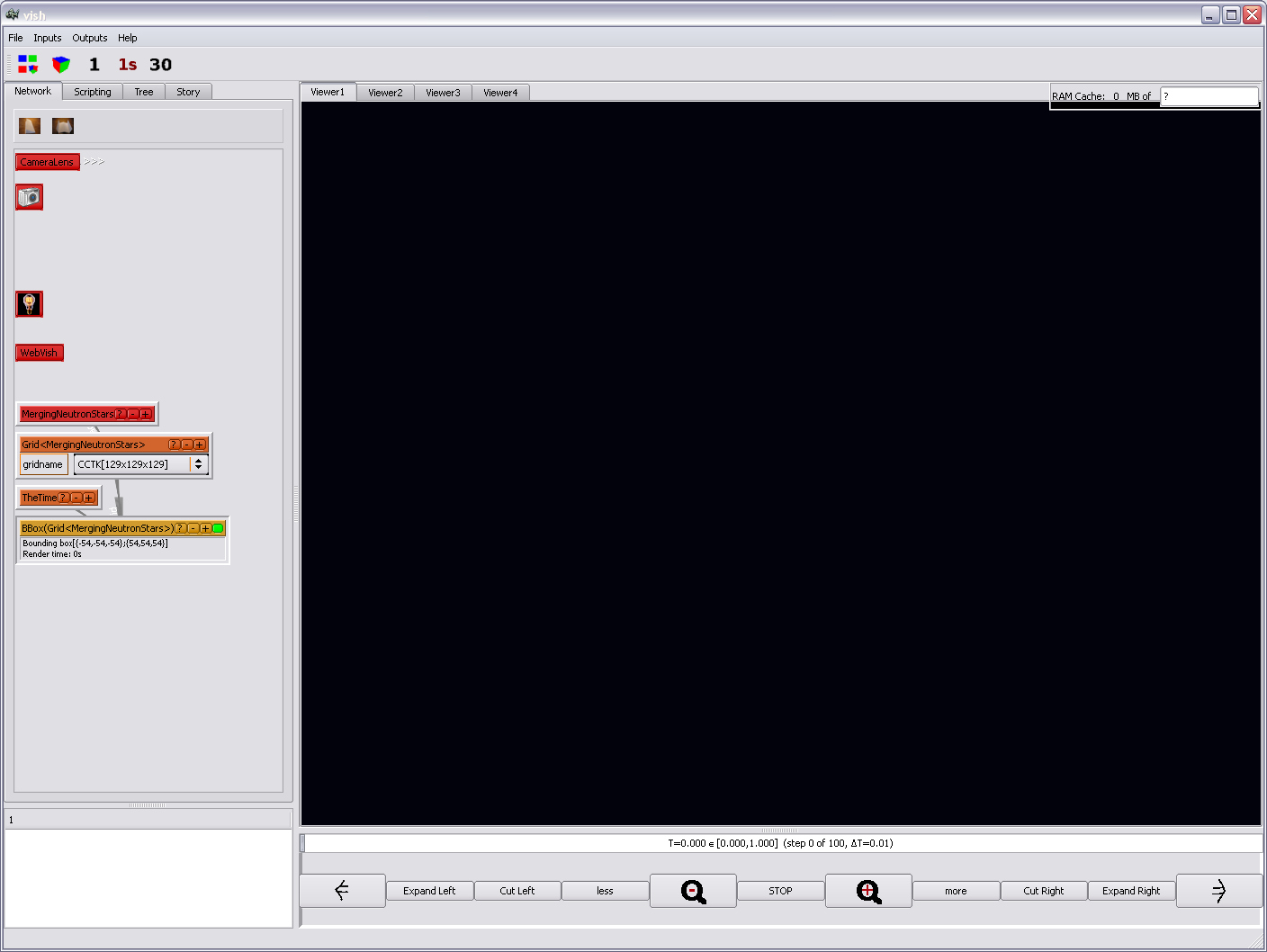
Unfortunately, you will not see a bounding box. Yet. This is just due to the
bounding box of this particular dataset being out of bounds of the viewing
volume. To adjust this discrepancy, click into the viewer on the right and
press the "space" bar on the keyboard. You should see the bounding box now:
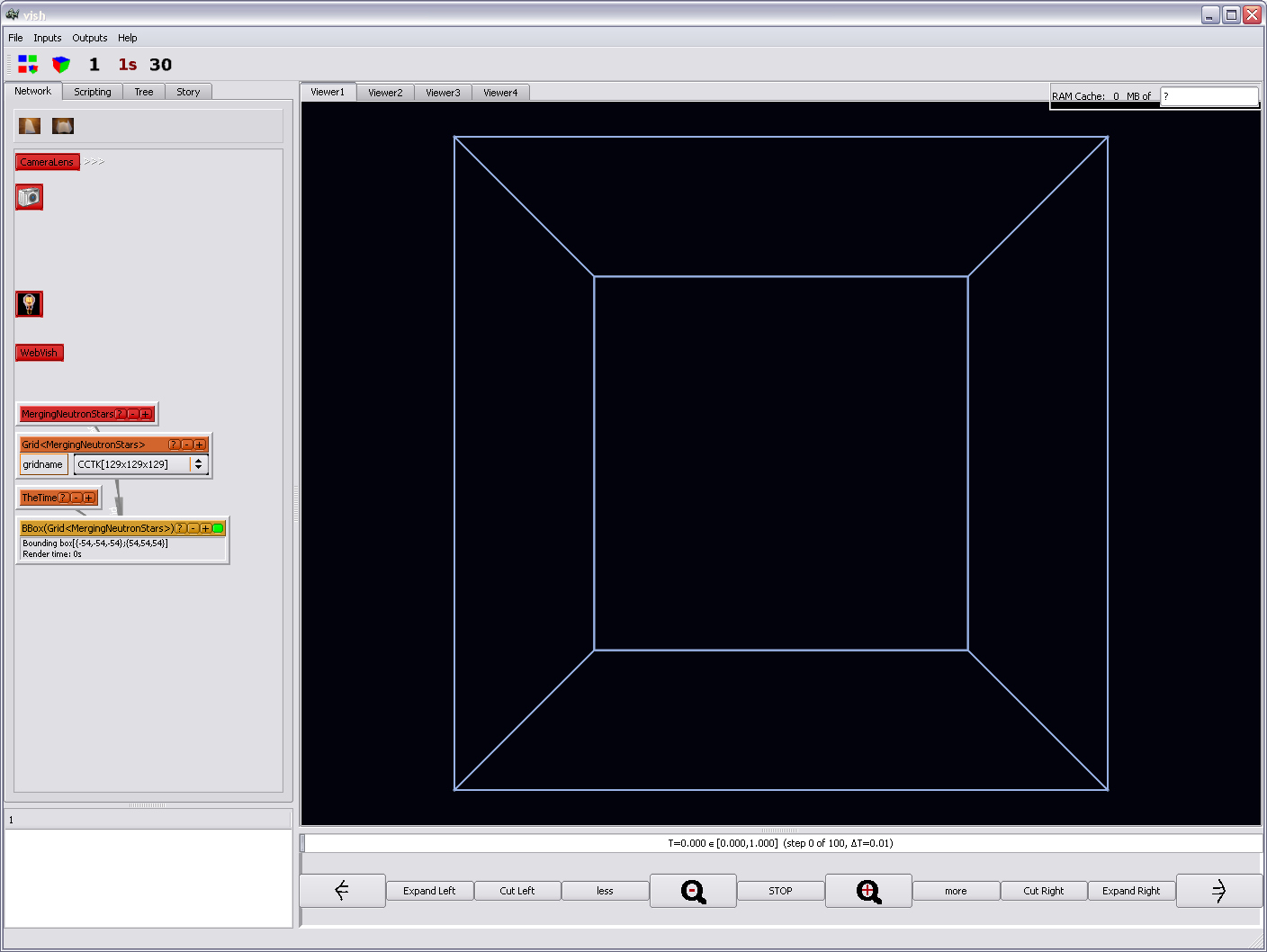
Remember this "space bar" adjustment whenever it might happen that nothing appears
in the viewer. Objects might just be outside of the viewing volume.
Once something is visible in the viewer, you can explore the mouse to spin around
the object and watch it from different sides. Left mouse rotates, right mouse
shifts, and mouse wheel or middle mouse moves forward/backwards.
Keyboard "X","Y","Z" and "x", "y","z" select coordinate-parallel views,
"t" rotates in the view plane, and also the cursor keys do something.
Should you ever feel the need for a different camera navigation,
source code for the navigation resides in vish/ocean/vcamera/VCameraNavigator.cpp .
If you just get lost in the view volume, pressing the space bar will adjust this mishap.
Now, for looking at the real data beyond just the bounding box, create a
Fiber/Scalarfield object from the orange Grid object.
There will be a yellow-orange box with one selection option, "rho" and "eps":
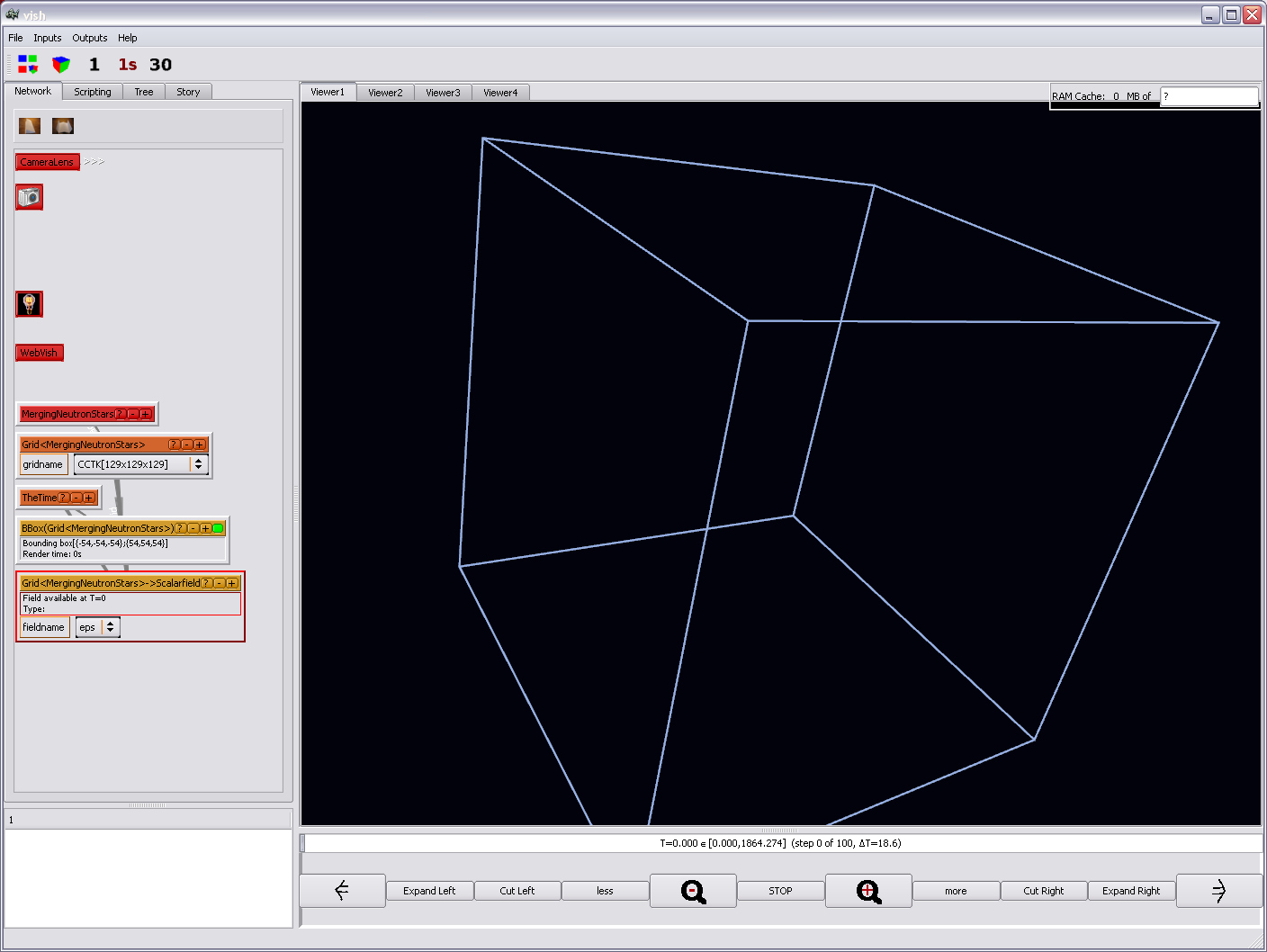
Now let us compute the numerical range of the data values in the "rho" field.
We will need this for a subsequent operation. So on the yellow-orange Scalarfield
object, add a Compute/Range object, which will - after a moment of
thinking - create a yellow icon, which contains the result of the operation:
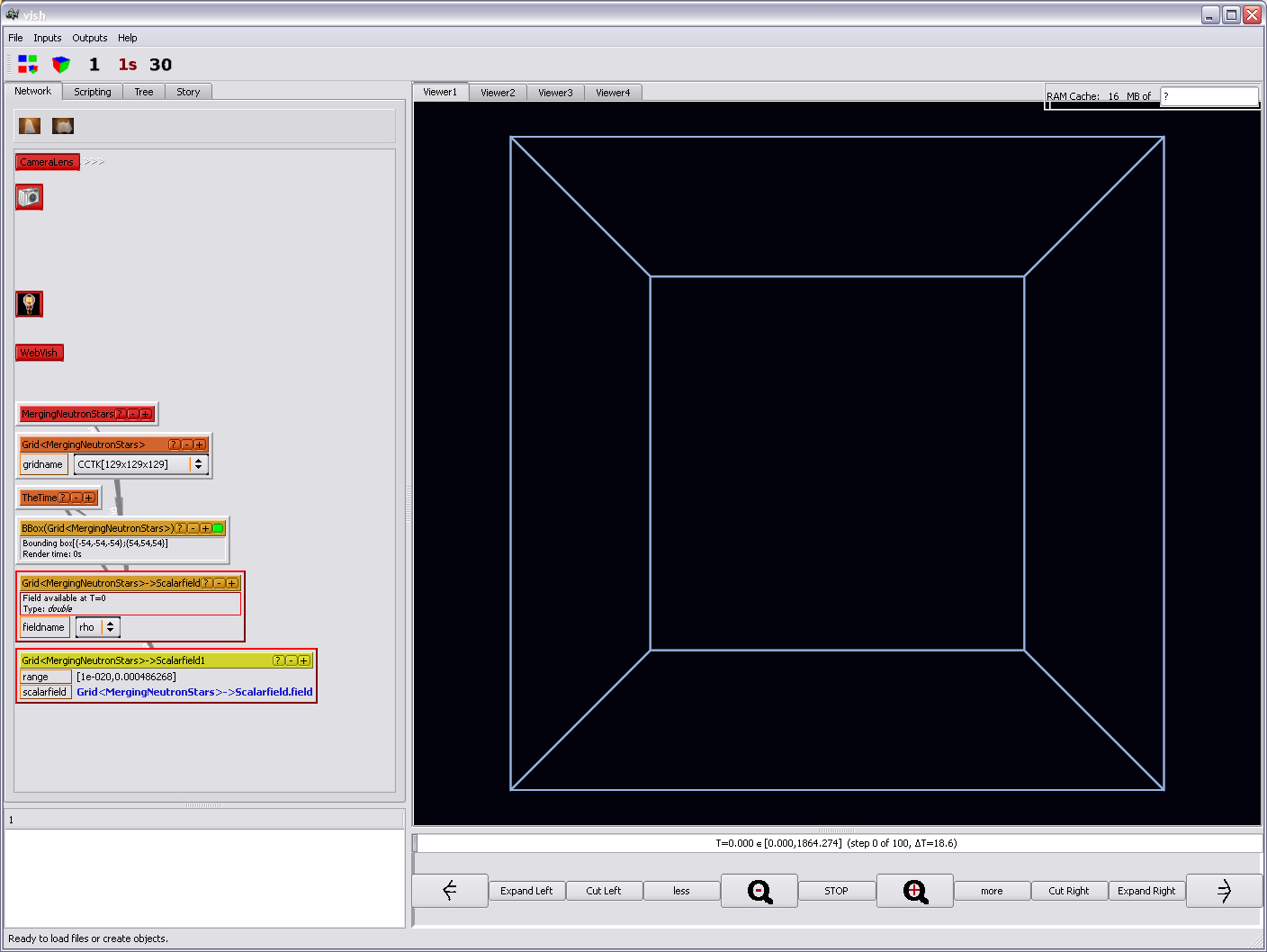
We want to attach an slice now with a colormap. It is possible to directly create
such output on the Scalarfield, but it will select an ugly colormap by default.
Thus rather lets create a prettier one in advance. To do so, right-click on
the gray area in the network tab, at any place which is not an icon.
From the menu, select the Colormaps/Colorramp object. It will
show up in red:
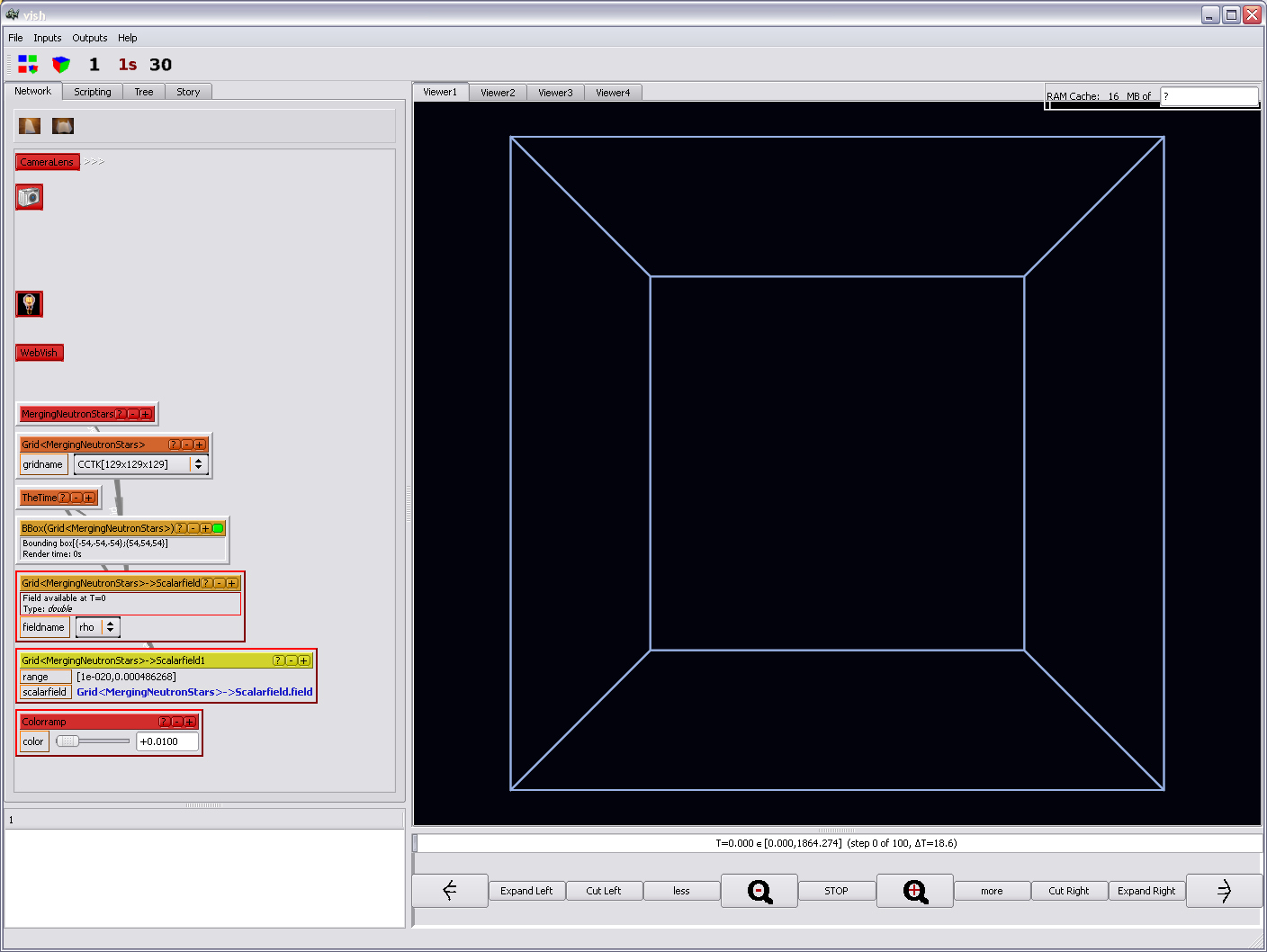
Now we are ready to attach a Display/Orthoslice to the yellow-orange
Scalarfield object. The new Orthoslice object will show up in yellow, and you
need to scroll down in the Network tab at the left to see all of its
parameters; also spin the 3D viewer such that you can see it a bit from
above:

Unfortunately, we do not see anything "data" yet. This is because the range
of the orthoslice is not adjusted with the range of the data.
For doing so, find the "datarange" parameter in the Orthoslice object,
and note that its value is a green text saying "Wizt::Range".
Please do not waste time with thinking what it means, but right-click
this text. You will get a context menu, with a selection of
"Createw new" entries, and one "Connect to" entry at the bottom of
the menu. Select this one. It means that the data range input of
the Orthoslice shall be connected with the result of the DataRange
computation as we had done before.
Now finally you will be able to see something:
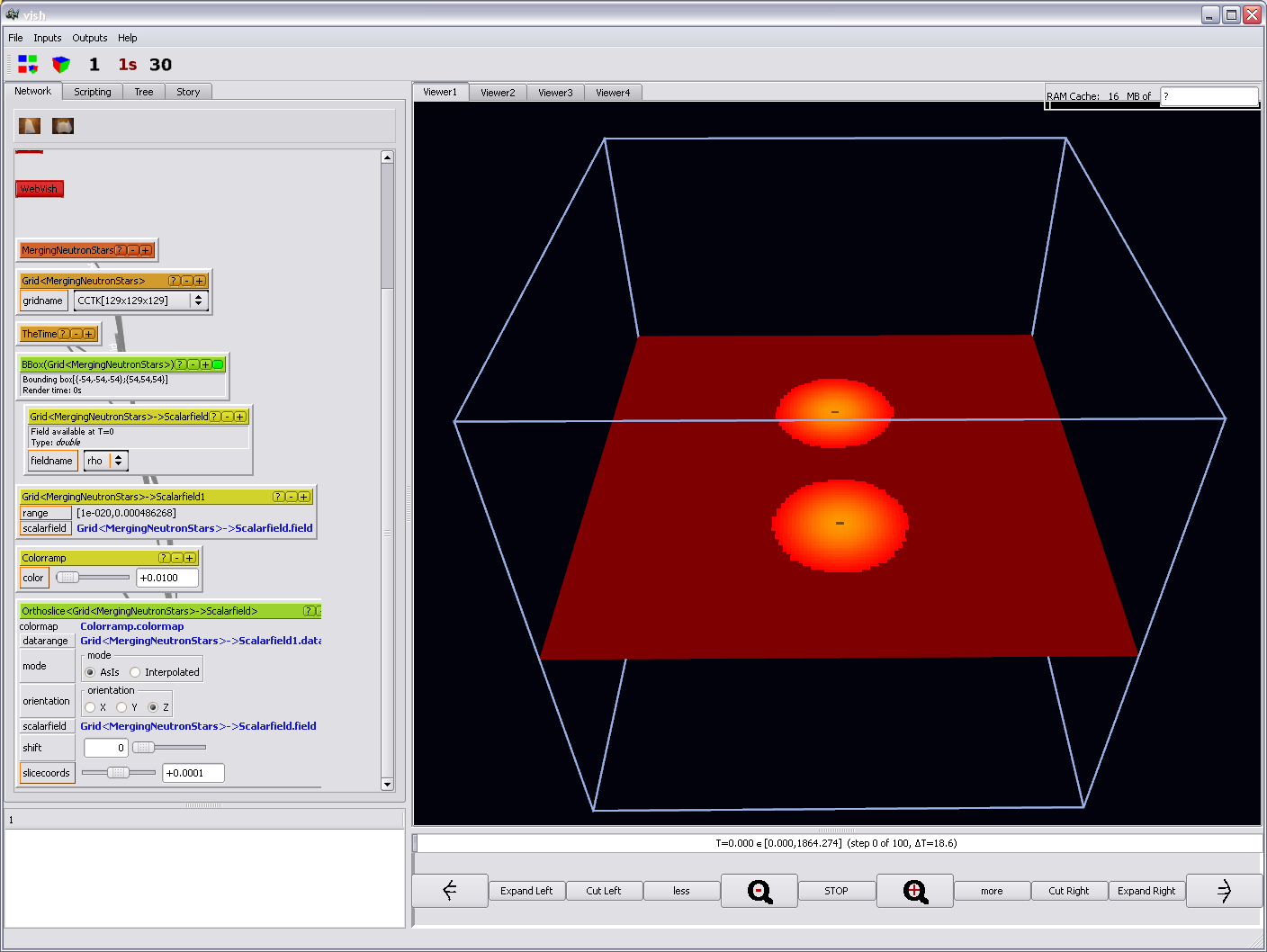
And the remaining visualization capabilities can be explored by playing with
the parameters, such as the time slider just below the viewer:
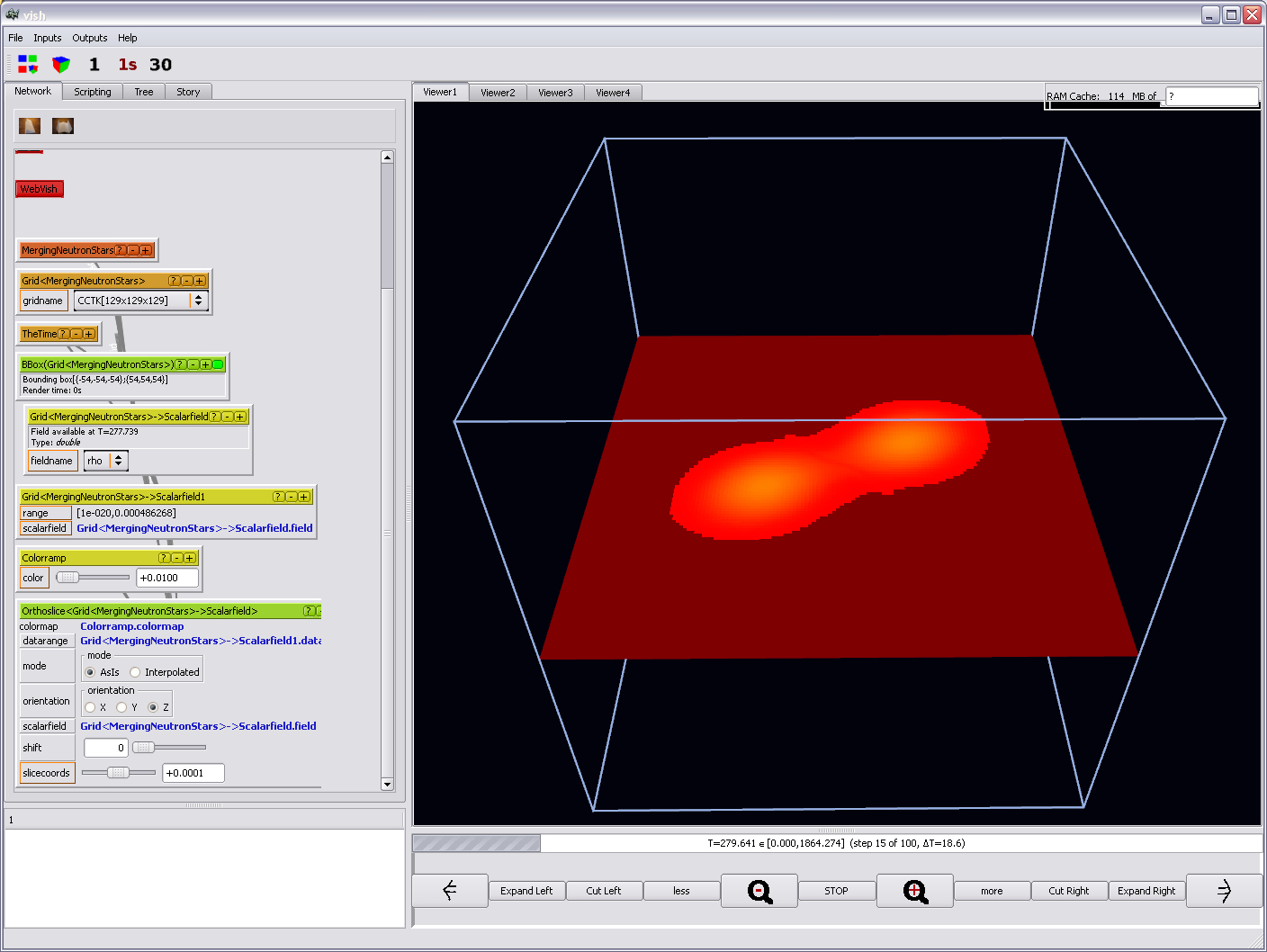

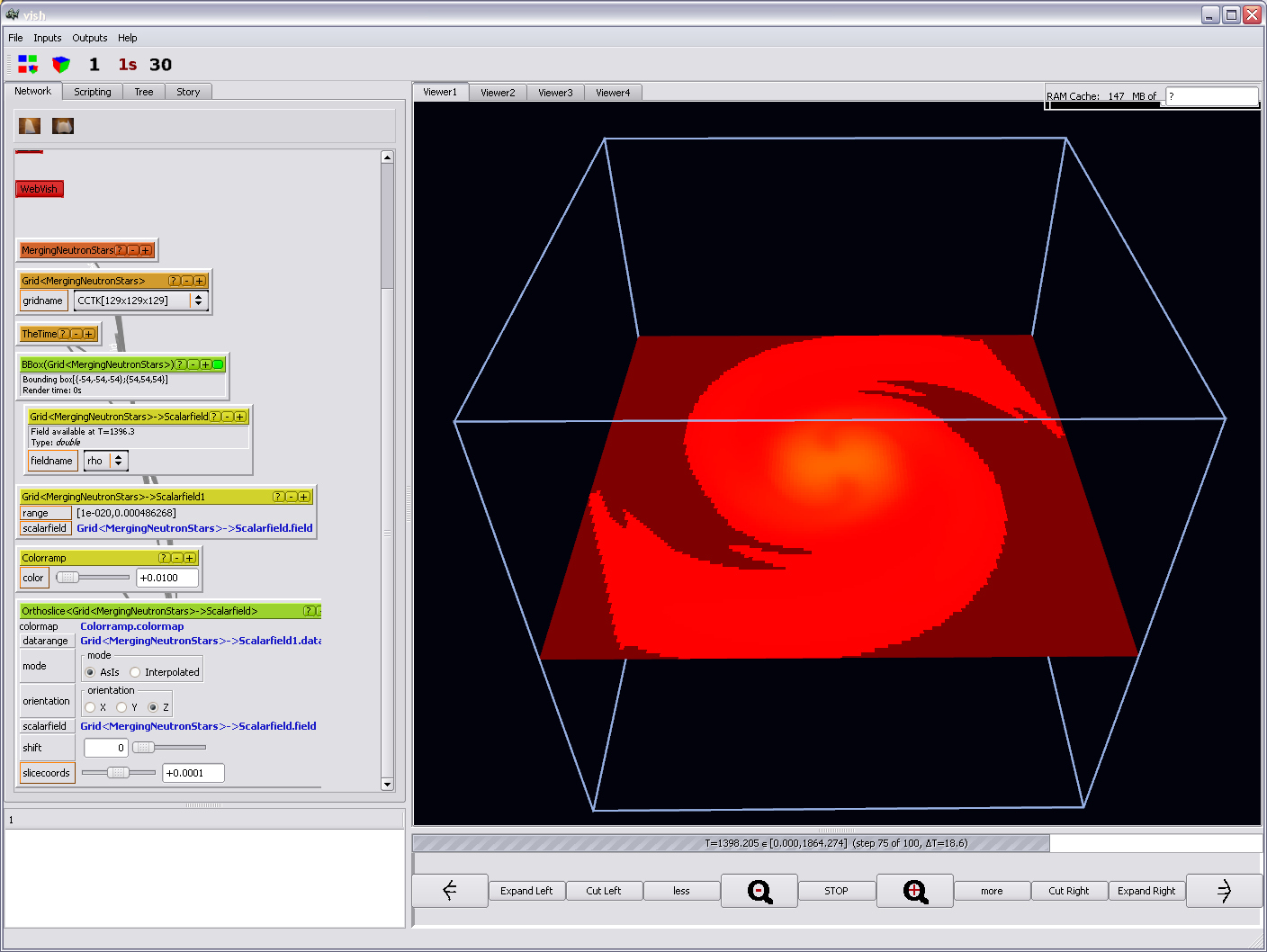
or the Colorramp's "color" parameter:

which actually looks much nicer once the Orthoslice interpolation is enabled,
but depends on the use case if that is desirable:
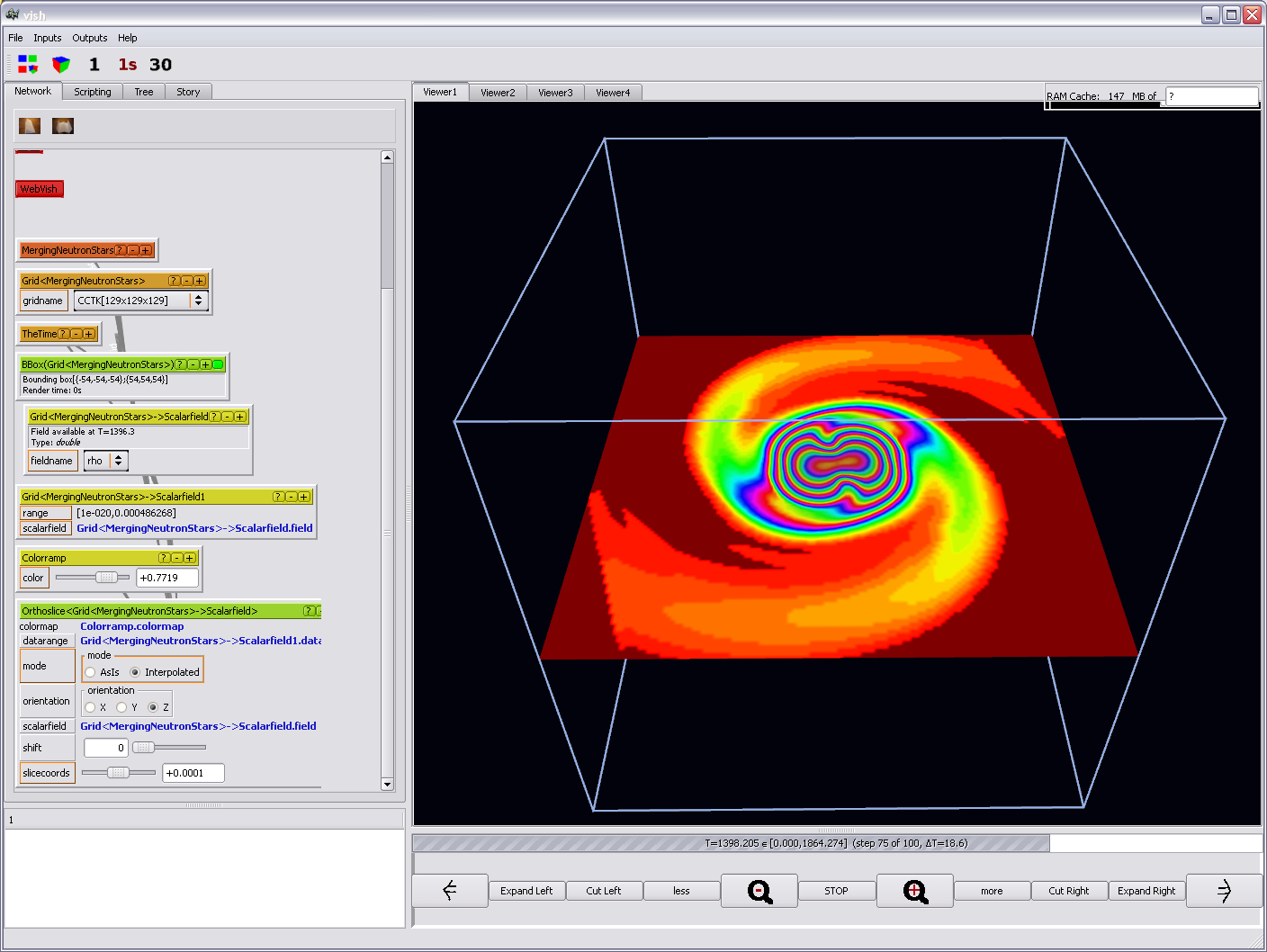
Other visualization options are invoked just similar and will be described below.
Creating a good visualization on a particular dataset requires careful
tuning of the available parameters anyway and requires experimenting
with the available options.
Creating Movies
For creating a movie, create a "Hollywood/VideoRecorder" object. Its parameters
are self-explaining, but it is recommended to not start with many frames for
the movie at first, since the dataset is huge and it will take a long time.
The VideoRecorder will create a number of files in the same directory
from where you invoked Vish.
Scripting
Instead of interactive usage, also a simple scripting functionality is supported by
Vish. The syntax is pretty close to the described interaction; best to get
started is to look at the .vis scripts that are provided with Vish.
They can be loaded by specifying on the command line when invoking Vish.
Scalar Fields
For scalar fields, the main Display options are
OrthoSlice, VolumeRender and IsoSurface.
Other display objects are mostly experimental.
Vector Fields
For vector fields, use the Display/VectorArrows or Display/VectorSpeckles
options for a direct display, or the Compute/WStreamlines and similar.
Tensor Fields
Direct visualization of 2nd order symmetric 3x3 tensor fields
is supported via the Display/TensorPatterns and Compute/EigenVectors
modules.
Adaptive Mesh Refinement
The script data/AMRortho.vis loads an example data set
demonstrating ortho slices on a dataset with mesh refinement.
Multipatch (curvilinear) Data and Particle sets
Are supported via Display/VertexSplats which can be used as a simple volume rendering for
scalar fields,
Display/VectorArrows and Display/VectorSpeckles as well as Compute/WStreamlines for
vector fields.
VolumeRender and Orthoslices are not supported for Multipatch data, as of now.














